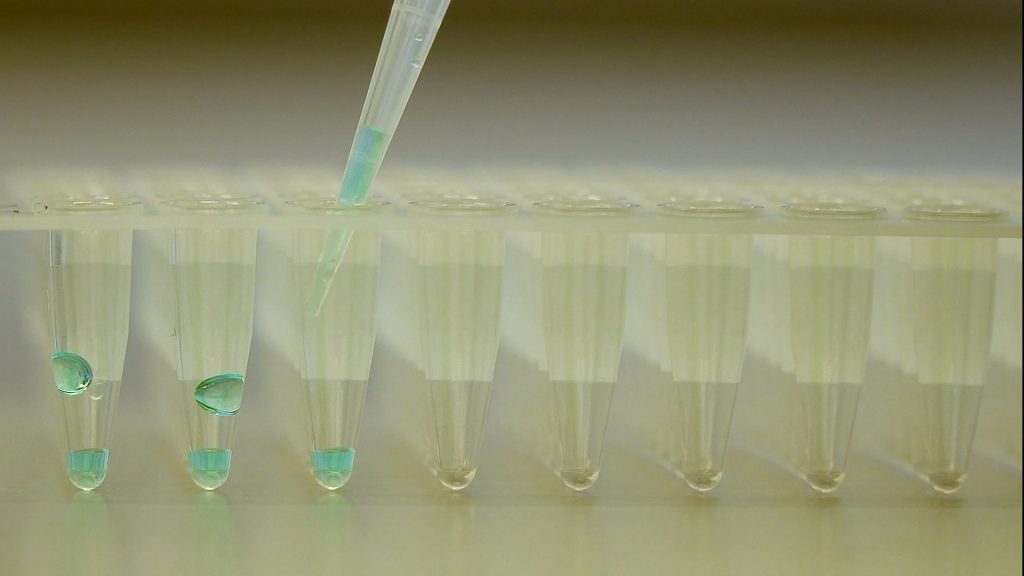
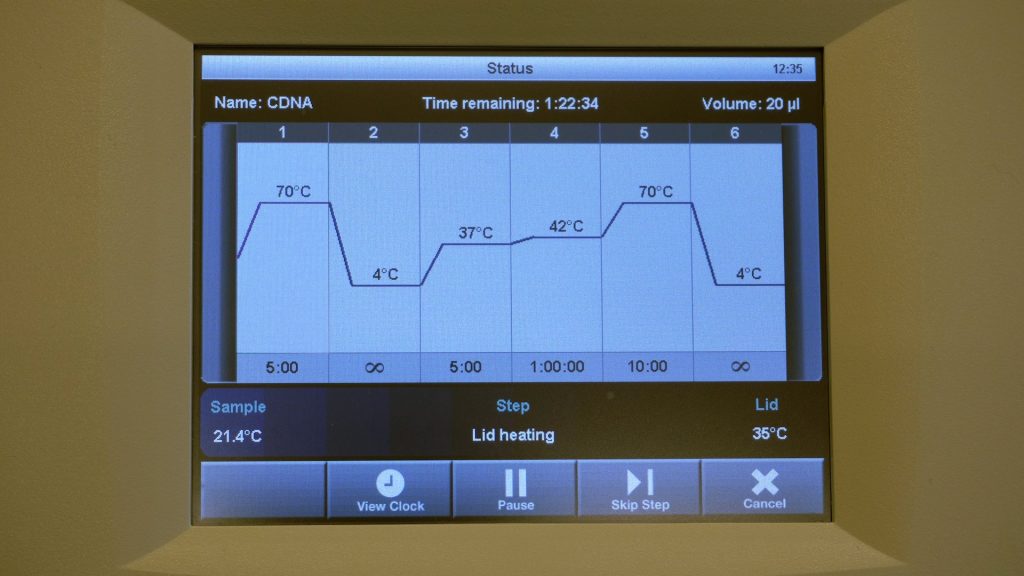
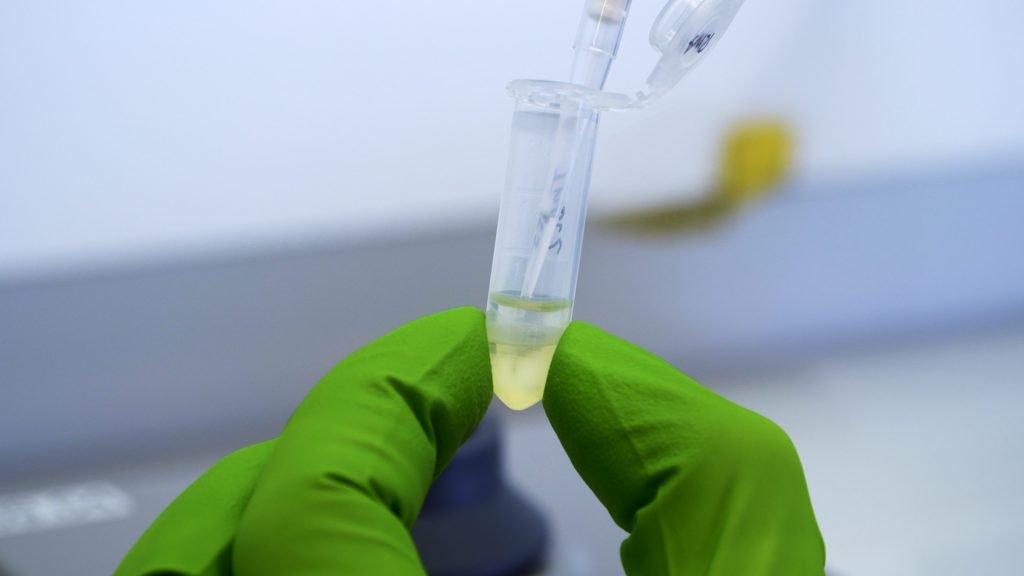
RNA isolation
This video shows how RNA isolation for qPCR works.
- Tineke Bijl
-
(0)
- 0 enrolled students
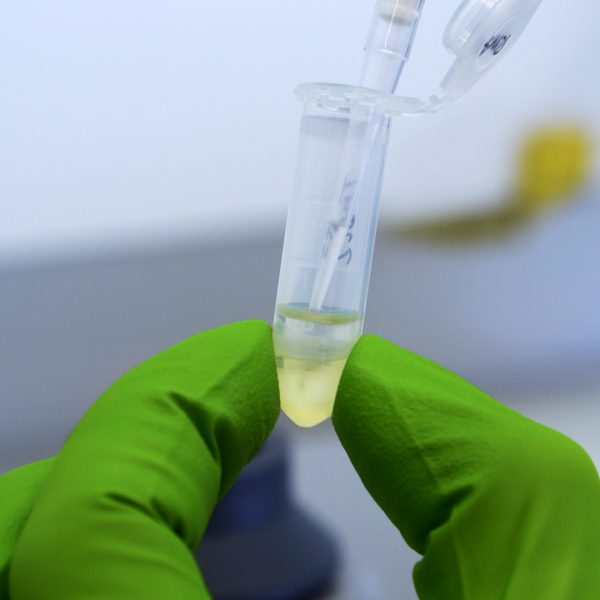
Description
Non-interactive video download link (mp4).
The script that has been used to create the video shown above can be downloaded as excel script file.
1 | Thaw the lysates on ice! |
2 | Put gloves on. Spray your bench 2x with RNase-Away / 2% SDS and clean the work surface with a tissue. |
3 | Work in the fume hood.
|
4 | Centrifuge the samples for 15 minutes at a speed of 12,000x g at 4 oC. |
5 | Very carefully transfer the top (aqueous) phase to a new RNase-free eppendorf tube. You continue with this eppendorf tube. Watch the video in the interactive potocol again for instructions. |
6 |
|
7 | Centrifuge the samples for 30 minutes at a speed of 12,000 x G at 4 oC. |
8 | Discard the supernatant. You can pat the eps upside down against a clean tissue to get the last bit of liquid out of the eppendrof tubes. |
9 | Add 100 µL RNase-free 75% EtOH to the samples. Vortex the samples briefly. |
10 | Centrifuge the samples at 15,000 x G for 5 minutes at 4oC. |
11 |
|
12 | You can work on the table from now on. Dissolve the samples in 20 µL RNase-free MilliQ. |
13 | The RNA concentrations of the samples are measured with the DS-11 spectrophotometer. As soon as 2-4 teams have finished their RNA isolation, an assistant will measure it for you. |
Which components does the supernatant of the RNA precipitation contain?
Feedback if incorrect: Think about the steps of the RNA isolation process. Which components are needed in the supernatant and which components need to be discarded? |
Select which operations are not required to maximize your RNA yield
Feedback if correct: Correct! Keep in mind that working in a fumehood does not change the amount of RNase. However this does not mean that you shouldn’t work in a fumehood!! You might use some hazardous chemicals in the protocol that are only to be used inside the fumehood. Ice is generally “dirty”. You don’t want to push the eppendorfs too deep into the ice, because they’ll get contaminated on the sides. You’ll definitely touch them to open the top and worse, you’ll probably touch the ice with your “clean” gloves. Your skin (and therefore your face) is full of RNase! Do not touch your face during the experiment. For a similar reason, don’t lean in closely to check something on your setup. It happens often that a hair (or eyelash, etc) end up in your experiment. Especially when working with PCR plates it is important to keep everything clean. Feedback if incorrect: Think of what is really necessary to work RNase-free. Which operations do we do for different reasons? |
The phase partition step in RNA isolation is often described using chloroform. For what reason is bromochloropropane (BCP) often used as an alternative for this step? a. BCP is used instead of chloroform because it is less toxic. b. Synthetic production of BCP is less complex and cheaper than chloroform. c. Chloroform is very susceptible to degradation over time, whereas BCP is not. d. The use of BCP decreases the possibility of contaminating RNA with DNA. e. Chloroform is colourless, whereas BCP is not. The phase partition is hard to observe with a colorless liquid. f. Generally speaking, BCP is most suitable for human tissue, whereas chloroform is more often used for other mammalian tissues. Feedback if correct: That is correct. Feedback if b: Synthetic production of BCP involves many steps and reacting agents and is in general terms more expensive to obtain compared to chloroform. Feedback if e: Both BCP and chloroform are colorless. Feedback if f: BCP usage does not hold restrictions on human tissue and is also commonly used for mammalian tissue. |
What should you do when there is two phases in your pipette tip after attempting to phase-separate the water phase from the organic phase? a. Repeat the centrifugation step.
b. Put the water phase back into the Eppendorf and pipet off the water phase again.
c. Taking some organic phase out together with the water phase is not a problem.
d. You need to start all over with the RNA isolation from the first step of the protocol.
|
How should the Eppendorf look after correct separation?
|
Download zip to import in LabBuddy.
Please make sure that your product exists and valid for this course
- Skill levelIntroduction video
- CategoryMolecular biology
Related videos
-
Free
qPCR
-
Free
cDNA synthesis
-
Free
RNA isolation
Copyright information
This video is created by Leiden Academic Centre for Drug Research (LACDR), Faculty of Science at Leiden University under a open Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. When using this video in its original version please refer to www.labprep.video. When adapting the video, mention the source ‘adapted based on the original version that is created by the labprep.video team’. It is not allowed to use the video for commercial purposes without consultation with the creators. You can contact us via info@labprep.video.